ORTHOSCOPE: instruction |
| Overview |
ORTHOSCOPE is a web tool to identify orthogroup members (orthologs and paralogs; see below) of a specific protein-coding gene of animals and plants. By uploading gene sequences of interest and selecting species genomes from more than 600 animals and plants, users can infer their functions and copy numbers according to gene-tree-based results. |
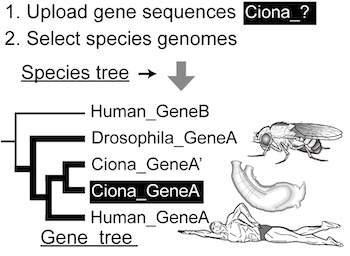 |
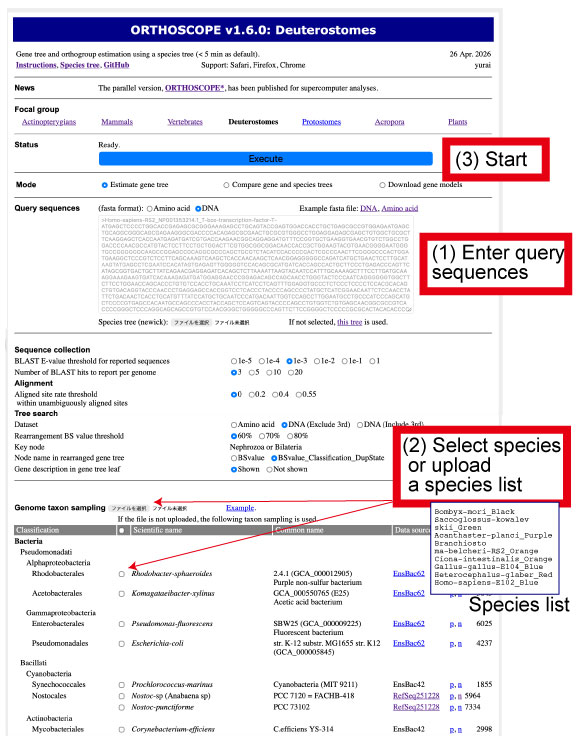
| Using sequences collected by BLAST search, ORTHOSCOPE estimates the gene tree, compares it with the species tree and identifies an orthogroup. ORTHOSCOPE works only for a specific gene and does not allow genome-scale analyses. A downloadable genome-wide pipeline is available at the ORTHOSCOPE* Github repository. |
|
| Quick Start |
| Further Reading |
 |
Japanese instruction (日本語の説明): Genome-wide analyses: |
| Orthogroup |
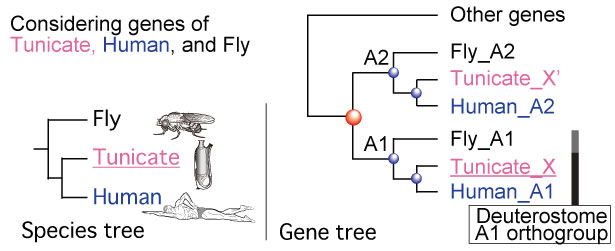
For identifying orthologs or genes with the same function, ORTHOSCOPE uses the concept of an orthogroup. We define an orthogroup as follows: a set of genes descended from a single gene in the last common ancestor of all the species being considered. This definition is similar to that of Emms and Kelly (2015), but differs in that we use the species tree to identify the common ancestor.
| Flow Chart |
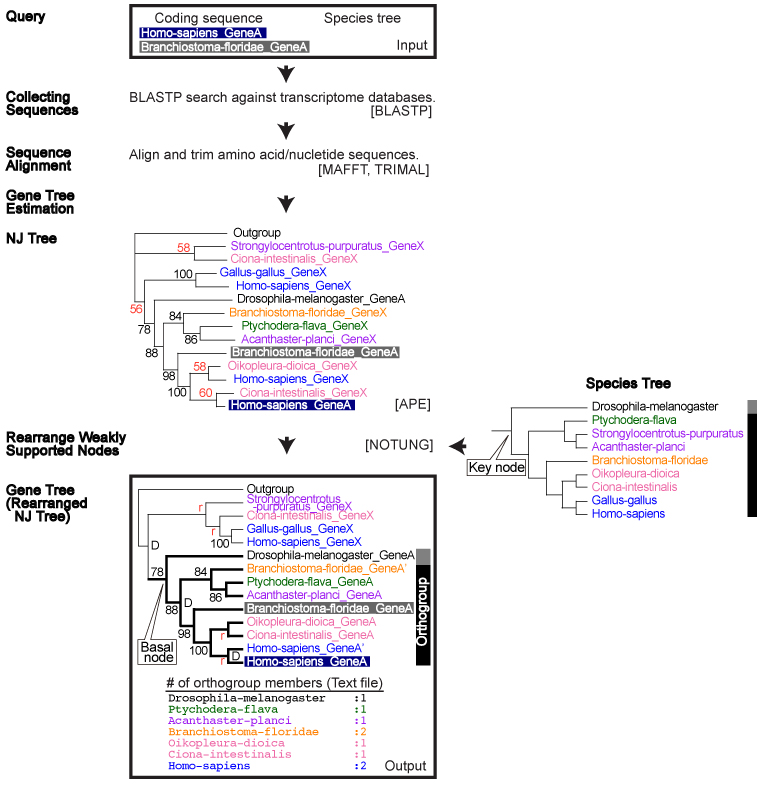
Dependencies:
BLAST v2.7.1+
MAFFT v7.356b
trimAl 1.2rev59
PAL2NAL v13
ape in R, Version5.0
FastME v2.0 for amino acid analyses
Notung-2.9
| Example Data |
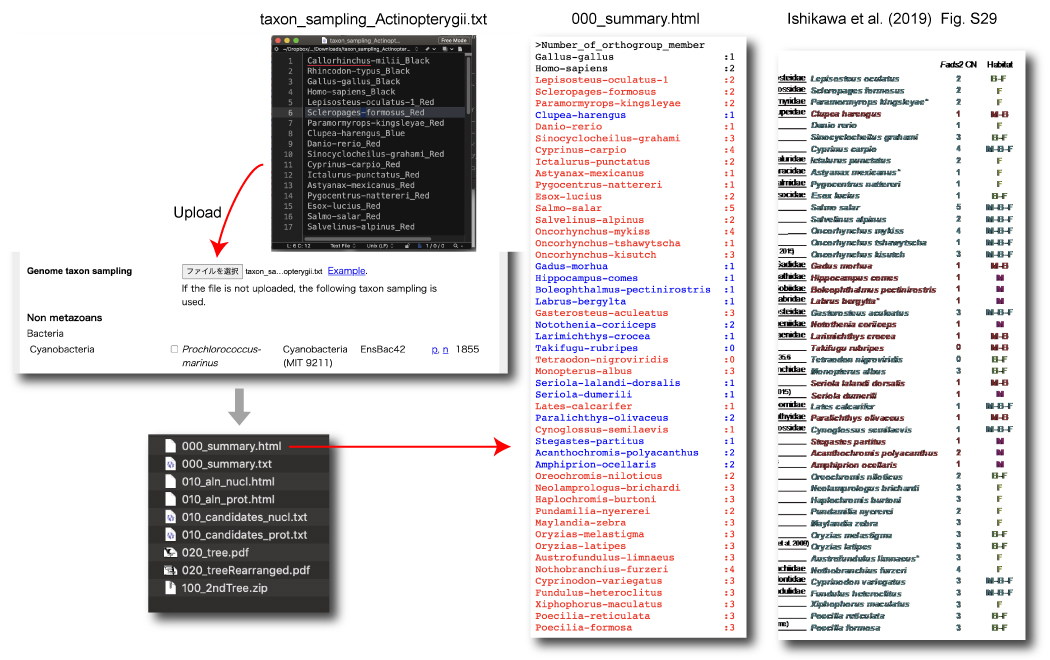
Ishikawa et al (2019)
Ishikawa, A, et al. 2019. A key metabolic gene for recurrent freshwater colonization and radiation in fishes. Science, 364: 886-9. Link.
Queries.
Taxon sampling.
To count Fads2 gene copies, these sequences were used for "Comparing gene and species trees" mode of "Focal group Actinopterygii".
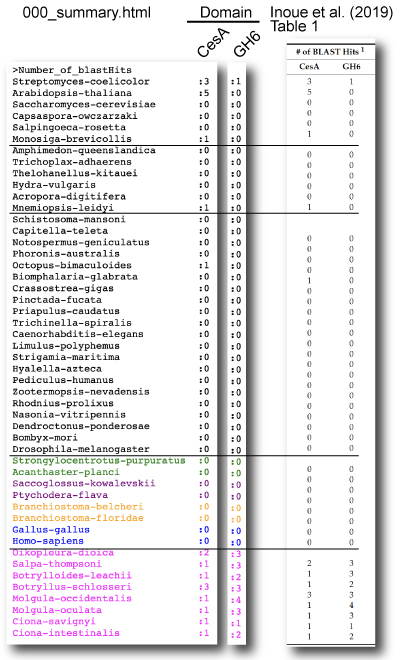
Inoue et al (2019)
| Inoue J, Nakashima K, and Satoh N. 2019. ORTHOSCOPE analysis reveals the presence of the cellulose synthase gene in all tunicate genomes but not in other animal genomes. Genes. 10: 294. Link Queries For queries, CesA gene sequences were separated into 2 parts: CesA (before TM7) and GH6 (after TM7) domains. Taxon sampling In this paper, maximum likelihood trees were estimated according to the process described in "Tree Estimation of Orthogroup Members". See below. |
 |
Inoue and Satoh (2019)
| Inoue J. and Satoh N. 2019. ORTHOSCOPE: an automatic web tool of analytical pipeline for ortholog identification using a species tree. 36:621–631. Link. |
|
Download query sequences from NCBI/Ensembl
From NCBI or Ensembl, query sequences can be downloaded.
For coding sequences, please select CDS as follows.
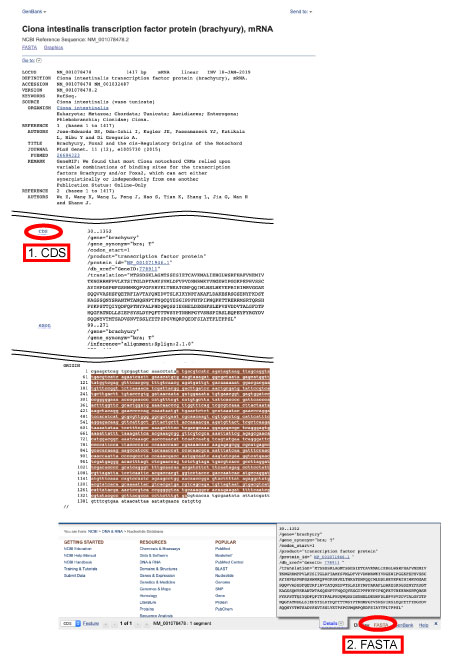

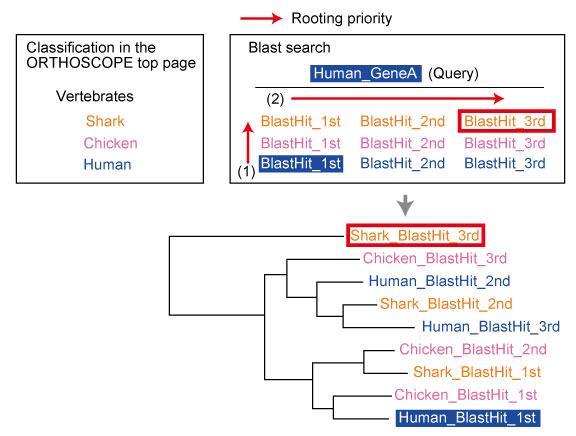
| Rooting Selection from BLAST Hits |
The sequence used for rooting the gene tree is determined by a two-step rule:
(1) Species selection: Among the species included in the analysis, the sequence from the species ranked highest in the species list on the top page.
(2) Sequence selection: Among sequences from the same gene model, the sequence most distantly related to the query sequence (or the topmost query sequence when multiple queries are provided), as determined by the lowest BLAST score.
| Tree Estimation of Orthogroup Members |
Analysis pipeline with example data: DeuterostomeBra_2ndAnalysis.zip
Installing dependencies
Estimation of the 2nd tree by the downloaded pipeline requires some dependencies to be installed and included in your system path.
IQ-TREE
Available here: https://iqtree.github.io
Install the latest version according to the above site.
Add the directory path to your PATH, for example:
Mafft v7.407 or laterexport PATH=$PATH:~/bin
Available here: https://mafft.cbrc.jp/alignment/software/ . After compilation, set your PATH following this site.
trimAl
Available here: http://trimal.cgenomics.org/downloads
Move into trimAl/source, type make, and then copy the executable:
pal2nal v14make cp trimal ~/bin
Available here: http://www.bork.embl.de/pal2nal/#Download
Change the permission of the Perl script and copy it:
APE in Rchmod 755 pal2nal.pl cp pal2nal.pl ~/bin
R (3.5.2) is available from CRAN. By installing R, rscript is installed automatically.
Install APE in R by typing the following command in the R console:
install.packages("ape")
Tree estimation
Using the downloaded pipeline, the second gene trees are estimated as follows:
- Based on the estimated rearranged NJ tree, users should manually select coding sequences of the orthogroup and outgroups. The pipeline then starts subsequent analyses.
- Selected sequences are aligned using MAFFT (Katoh et al. 2005).
- Multiple sequence alignments are trimmed using trimAl (Capella-Gutierrez et al. 2009) with the option gappyout.
- Corresponding cDNA sequences are forced onto the amino acid alignment using PAL2NAL (Suyama et al. 2006).
- Phylogenetic analysis is performed using IQ-TREE 2 (Minh et al. 2020),
The actual process is as follows:
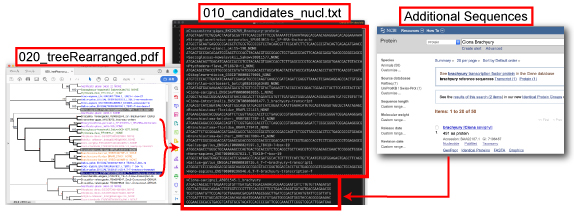
- Decompress DeuterostomeBra_2ndAnalysis.zip, open the DeuterostomeBra_2ndAnalysis directory, and extract 100_2ndTree.tar.gz.
- Select an appropriate outgroup and orthogroup members and save them as 010_candidates_nucl.txt. The outgroup sequence should be placed at the top of the alignment. Additional sequences can be included.
- Change directory to 100_2ndTree.
- Run the pipeline:
./100_estimate2ndTree.py
- The ML tree is automatically saved.
| Collecting Query Sequences from an Assembly Database |
Here, we demonstrate this procedure using vertebrate ALDH1A and actinopterygian PLCB1 (Inoue and Satoh, 2019).
- Download Coregonus lavaretus TSA file (GFIG00000000.1) from NCBI.
- Translate raw sequences into amino acid and coding sequences using TransDecoder.
./TransDecoder.LongOrfs -t GFIG01.1.fsa_nt
- Create blast databases using BLAST+.
makeblastdb -in longest_orfs.pep -dbtype prot -parse_seqids makeblastdb -in longest_orfs.cds -dbtype nucl -parse_seqids
- Perform a BLASTP search against amino acid database.
blastp -query query.txt -db longest_orfs.pep -num_alignments 10 -evalue 1e-12 -out 010_out.txt
- Retrieve BLAST top hit sequences from coding sequence file using their sequence IDs.
blastdbcmd -db longest_orfs.cds -dbtype nucl -entry_batch queryIDs.txt -out 020_out.txt
| History |
| Date | Version | Revision |
|---|---|---|
| 27 Oct 2025 | Version 1.6.0 | Released. An alignment overview was newly added. New data were added for the following taxa: Cyanobacteria (Nostoc-sp, Nostoc-punctiforme), Chlorarachniophyta (Bigelowiella-natans), Oomycota (Bremia-lactucae), Bacillariophyta (Phaeodactylum-tricornutum, Thalassiosira-pseudonana), Ciliophora (Paramecium-tetraurelia), Colpodellida (scaffold-Chromera-velia), Apicomplexa (Toxoplasma-gondii), Haptophyta (scaffold-Pavlovales-sp, chr-Rebecca-billardiae, scaffold-Diacronema-lutheri, contig-Prymnesium-sp, chr-Prymnesium-parvum, chr-Isochrysis-galbana, Emiliania-huxleyi-E62), Euglenozoa (chr-Euglena-gracilis), Cryptophyta (Guillardia-theta), Rhodophyta (Chondrus-crispus-RS, Galdieria-sulphuraria-RS, Cyanidioschyzon-merolae-RS, contig-Cyanophora-paradoxa), Chlorophyta (Ostreococcus-tauri, Micromonas-commoda, scaffold-Coccomyxa-elongata, scaffold-Coccomyxa-viridis, chr-Coccomyxa-sp, Coccomyxa-subellipsoidea-T, Coccomyxa-subellipsoidea, Auxenochlorella-protothecoides, scaffold-Parachlorella-kessleri, Chlorella-variabilis, Sphaeropleales, Chlamydomonadales, Chlamydomonas-reinhardtii, scaffold-Chlamydomonas-eustigma, Caulerpa-lentillifera), Chlorokybophyceae (scaffold-Chlorokybus-atmophyticus), Mesostigmatophyceae (scaffold-Mesostigma-viride), Klebsormidiophyceae (scaffold-Klebsormidium-nitens), Urochordata (Salpa-thompsoni-t, Ciona-intestinalis-E115). |
| 25 Feb 2025 | New data added: A close relative of zebrafish (Danio aesculapii). | |
| 3 Dec 2025 | New data added: Fungi (Trametes versicolor, Phanerochaete chrysosporium). | |
| 28 Nov 2025 | New data added: Phaeophyceae (brown algae) (Cladosiphon-okamuranus-K, -O, -C). | |
| 26 Nov 2025 | New data added: Symbiodiniaceae (Durusdinium trenchii), Phaeophyceae (brown algae) (Ectocarpus-siliculosus, Nemacystus-decipiens, Cladosiphon-okamuranus), Rhodophyta (red algae) (Chondrus-crispus, Galdieria-sulphuraria, Cyanidioschyzon-merolae), Viridiplantae (Chlamydomonas-reinhardtii), Bacteria (Rhodobacter-sphaeroides, Komagataeibacter-xylinus, Pseudomonas-fluorescens, Escherichia-coli). | |
| 24 Nov 2025 | New data added: Zebrafish (Danio-rerio-Ens115 and Danio-rerio-RS). | |
| 23 Nov 2025 | New data added: Green bottle fly, Lucilia-sericata. | |
| 5 Nov 2025 | New data added: Cephalochordata (Branchiostoma-belcheri-R3, Branchiostoma-floridae-RS3). | |
| 4 Nov 2025 | New data added: Hemichordata (Saccoglossus-kowalevskii-E), Echinodermata (Anneissia-japonica-E, Asterias-rubens-E, Acanthaster-planci-E, Strongylocentrotus-purpuratus-E62), Cephalochordata (Branchiostoma-lanceolatum-kl), Vertebrata (Gallus-gallus-Ens115, Homo-sapiens-Ens115). | |
| 28 Oct 2025 | New data added: Phaeophyceae (brown algae) (Ectocarpus-siliculosus, Nemacystus-decipiens, and Cladosiphon-okamuranus) and Rhodophyta (red algae) (Chondrus-crispus, Galdieria-sulphuraria, and Cyanidioschyzon-merolae). | |
| 27 Oct 2025 | New data added: Dinoflagellate (Durusdinium-trenchii) and Fungi (Chaetomium-globosum, Neurospora-crassa, Trichoderma-reesei, Batrachochytrium-dendrobatidis). | |
| 19 Oct 2025 | Version 1.5.8 | Released. New data added: Tunicate, Bacteria (Kitasatospora-setae and Streptomyces-griseus), and Viridiplantae (Chlamydomonas-reinhardtii). |
| 19 May 2025 | Version 1.5.7 | Released. The updated data of a tunicate, Clavelina-lepadiformis-SU, were added. |
| 21 Jan 2025 | Bootstrap analyses are started with the same seed number by the R script, set.seed(123). | |
| 28 Dec 2024 | Version 1.5.6 is released. (1) trimal1.5 is used to handle stop codons (*) in amino acid sequence (e.g. Homo-sapiens-RS2_NP036380.2). (2) In rearranged trees, detailed node labels are shown. (3) Gene description can be deleted. | |
| 2 Oct 2024 | Version 1.5.5 is released. | |
| 24 July 2024 | Updated data of an appendicularian (Oikopleura-dioica-OKI2018) were newly added. | |
| 4 July 2024 | Updated data of a tunicate (Ciona intestinalis KY21) were newly added. | |
| 6 June 2024 | Version 1.5.3 | Released. The "Downloading gene models" mode was newly constructed. |
| 1 May 2024 | Data of two tunicates (Halocynthia roretzi and H. aurantium) were newly added. | |
| 26 Mar 2024 | Data of two sharks (Mobula-hypostoma and Hemiscyllium-ocellatum) were newly added. | |
| 11 Mar 2024 | Data of an Echinoderm (Lytechinus-pictus) were newly added. Updated versions of two vertebrates (Homo sapiens and Gallus gallus) data were added. | |
| 3 Mar 2024 | New version data of C. elegans and D. melanogaster were newly added. | |
| 29 Feb. 2024 | Data of three flowering plants (Phragmites-australis, Carica-papaya, and Raphanus-sativus) were newly added. | |
| 14 Feb. 2024 | Data of seven eudicot plants (Eucalyptus grandis, Pistacia vera, Gossypium raimondii, Arabis alpina, Eutrema salsugineum, Brassica rapa, and Arabidopsis lyrata) were newly added. | |
| 9 Feb. 2024 | Data of five anthozoans (Dendronephthya-gigantea, Actinia-tenebrosa, Xenia-sp, Exaiptasia-diaphana, and Pocillopora-verrucosa) were newly added. | |
| 3 Feb. 2024 | Data of seven bivalves (Mercenaria-mercenaria, Pecten-maximus, Ylistrum-balloti, Mytilus-californianus, Saccostrea-echinata, Ostrea-edulis, and Crassostrea-angulata) were newly added. | |
| 1 Feb. 2024 | Data of seven teleosts (Coregonus clupeaformis, Salmo trutta, Salvelinus namaycush, Oncorhynchus nerka, Oncorhynchus gorbuscha, Oncorhynchus keta, and Cololabis saira) were newly added. | |
| 31 Jan. 2024 | Data of a pearl oyster (Chromosome-scale, haplotype-phased genome assembly A and B) were newly added. | |
| 28 Jan. 2024 | Data of a conger eel (Conger conger) and a conifer (Cryptomeria japonica) were newly added. | |
| 24 Aug. 2023 | Data of a ray-finned fish (Amia calva) were newly added. | |
| 23 Apr. 2023 | Data of a coral (Pocillopora damicornis) were newly added. | |
| 21 Jan. 2023 | Data of two ferns (Marsilea vestita and Ceratopteris richardii) were newly added. | |
| 18 Dec. 2022 | Data of a sawfish (Pristis pectinata) were newly added. | |
| 28 Aug. 2022 | Data of Maidenhair tree (Ginkgo biloba), two ferns (Azolla filiculoides and Salvinia cucullata), and gymnosperms (Ginkgo biloba, Cycas-micholitzii, Gnetum-montanum, Taxus-baccata, Pseudotsuga-menziesii, Picea-glauca, and Pinus sylvestris) were newly added. | |
| 9 Aug. 2022 | Data of a hornwort (Anthoceros angustus), liverworts (Marchantia polymorpha, Male Tak1 and male and female) were newly added. | |
| 9 Aug. 2022 | Data of a basal streptophyte alga (Penium-margaritaceum) were newly added. | |
| 2 Jul. 2022 | Data of two plants (Leersia perrieri and Camelina sativa) were newly added. | |
| 2 Jul. 2022 | Data of two sharks (C. plagiosum and S. fasciatum) and 5 plants (Brachypodium-distachyon, Hordeum-vulgare, Secale-cereale, Aegilops-tauschii, Triticum-spelta, and Actinidia-chinensis) were newly added. | |
| 9 Jun. 2022 | Data of a sweet orange were newly added. | |
| 22 May 2022 | Version 1.5.2 | Bug fix release: In result pages derived from "Comparing gene and species trees" mode, query replacements are shown. Amino acid analyses were fixed by limiting character numbers of name lines up to 60. |
| 19 May 2022 | New mirror site, AORI:viento, is added. | |
| 18 Dec. 2021 | Data of a sea urchin (Lytechinus) and 3 crustaceans (Penaeus japonicus, Homarus, and Portunus) were newly added. | |
| 11 Dec. 2021 | Data of 2 ancient fishes (Polypterus, Polyodon) were newly added. | |
| 14 Aug. 2021 | Data of 4 liliopsid data (e.g., Dioscorea, Asparagus, Zingiber, and Ananas) were newly added. | |
| 13 Aug. 2021 | Data of 7 fabales (e.g., Glycine-max) were newly added. | |
| 25 Jul. 2021 | Data of Carcharodon carcharias (Great white shark) was newly added. | |
| 21 Jul. 2021 | Data of Caulerpa lentillifera (Siphonous green alga) was newly added. | |
| 19 Jun. 2021 | Data of seven plants (EnsPlant51) were newly added. | |
| 10 Apr. 2021 | Version 1.5.1 | Released. A focal group, Plants, was newly added. |
| 11 Feb. 2021 | Data of a shrimp (Penaeus monodon) and a tunicate (Styela clava) were newly added. | |
| 6 Feb. 2021 | Data of 2 sharks (Callorhinchus-milii-E102 and Scyliorhinus-canicula) and 18 teleosts (e.g. Hucho-hucho) were newly added. | |
| 5 Feb. 2021 | Data of 6 Oryzias individuals (Oryzias melastigma E102, O. javanicas, O. sinensis, O. latipes HSOK, HNI, HdrR E102) were newly added. | |
| 1 Feb. 2021 | Data of 2 asteroids (Asterias rubens and Patiria miniata [RefSeq]) and a lancelet (Branchiostoma lanceolatum [Ensembl]) were newly added. | |
| 29 Dec. 2020 | From ver. 1.5.0, each gene model can be downloaded by clicking p (amino acid sequence) or n (coding sequence) at the right of each species line. | |
| 29 Dec. 2020 | Version 1.5.0 | Text areas were introduced for sequence uploading. In conjunction with the renewal, the file uploading system was closed. |
| 24 Dec. 2020 | Gene model data were newly added for 4 snakes (Pantherophis guttatus, Thamnophis elegans, Naja naja, and Laticauda laticaudata). | |
| 6 Dec. 2020 | Version 1.2.2 | Gene model data were newly added for 3 sharks (Scyliorhinus torazame, Chiloscyllium punctatum, and Rhincodon typus), human (Homo sapiens Ens102), and chicken (Gallus gallus Ens102). |
| 6 Sep. 2020 | Version 1.2.1 | Gene model data were newly added for an echinoderm (Anneissia japonica) and replaced with TSA data for an acoela (Hofstenia-miamia). |
| 30 Aug. 2020 | Data of Sterlet (Acipenser ruthenus) and European eel (Anguilla anguilla) were newly added. | |
| 1 Jun. 2020 | Version 1.2.0 | Released. A focal group, Acropora, was newly added. |
| 1 Jun. 2020 | Version 1.1.0 | Released. Data of Amblyraja radiata (Thorny skate) was newly added. |
| 14 Jan. 2020 | The batch uploading was implemented for taxon sampling. <img src="images/batch_tsampling.jpg"> | |
| 6 Nov. 2019 | ORTHOSCOPE-Mammalia was newly created and data of 46 mammals were newly added. | |
| 6 Oct. 2019 | Gene model databases (fasta files of amino acid and coding sequences) can be downloaded from zenodo (10.5281/zenodo.2553737). | |
| 2 Oct. 2019 | Data of Pacific white shrimp (Penaeus vannamei) were newly added. | |
| 5 Sep. 2019 | Data of 2 molluscs (Octopus vulgaris, Pomacea canaliculata) were newly added. | |
| 21 Aug. 2019 | Column of Seqs (# of sequence in each gene model) was added. | |
| 21 Aug. 2019 | Data of 6 actinopterygians (Erpetoichthys calabaricus, Denticeps clupeoides, Carassius auratus, Electrophorus electricus, Tachysurus fulvidraco, Pangasianodon hypophthalmus), 2 amphibians (Rhinatrema bivittatum, Microcaecilia unicolor), and 3 lepidosaurians (Notechis scutatus, Podarcis muralis, Pseudonaja textilis) were newly added. | |
| 19 Apr. 2019 | Negative branch lengths are replaced with 0 in the tree drawing (R script). Gene_tree$edge.length[Gene_tree$edge.length<0]<-0 | |
| 25 Jan. 2019 | Version 1.0.2 | Released. For Inoue et al. 2019, Data of Archaea, Plants, Bacteria, and Urochordata were newly added. |
| 21 Dec. 2018 | Version 1.0.1 | Released. In the rearranged gene tree, nodes identified as speciation events were marked with "D". |
| 18 Dec. 2018 | Version 1.0.1.beta | Xenacoelomorph, platyhelminth, priapulid, avian data were newly added. |
| 10 July 2018 | Version 1.0 | Published in Inoue and Satoh (2018). |
| Citation |
Inoue J, Satoh N. 2019. ORTHOSCOPE: An automatic web tool for phylogenetically inferring bilaterian orthogroups with user-selected taxa. Molecular Biology and Evolution 36:621–631. Link.